Biomacromolecules
The Machinery of Life
DNA, RNA, and proteins are all biomacromolecules that act as sophisticated molecular machines in the cells. The growth, division, and apoptosis of a cell rely on the harmonic operation of millions of such molecular machines.
The double helical structure of DNA proposed by Watson and Crick in 1953 and the three-dimensional structure of myoglobin solved by Kendrew et al. marked the beginning of a molecular-level understanding of how these molecular machines work. The primary purpose of structural biology, which was established afterwards, is to study the function of biomacromolecules in the cell based on their three-dimensional structures. The structure and the function of proteins are the main focus of structural biologists. Scientists have solved more than 100,000 structures of proteins by the means of X-Ran diffraction, electron microscopy, and NMR by 2015. But most of these structures are similar. If we categorize these proteins by their similarities, approximate 2,000 big protein families can be obtained. Scientist now can predict a few simple protein structures by computer simulation according to these known structures. However, there is no computer algorithm that can successfully predict any protein structure just from the sequence the ammonia acids without any reliance on the established structures. In addition, scientists have discovered that approximately 40% of proteins do not have a specific structure as myoglobin. Such proteins are called naturally disordered proteins, some of which will form specific structures when they combine with other biomacromolecules or small molecules. How to predict the three-dimensional structure of proteins from the ammonia acid sequence and the study of biological function of naturally disordered proteins are the two primary challenges for structural biologists.
Computer graphics was in a primitive stage at the early period of structural biology (1950-1970). To spread this novel but fascinating subject to public, an American artist Irving Geis made outstanding contributions. The DNA and proteins paintings he created greatly influenced a generation of structural biologists and became the foundation of representing biomacromolecules by computer graphics. We redraw three of Geis’ masterpieces in this book to salute this great artist.
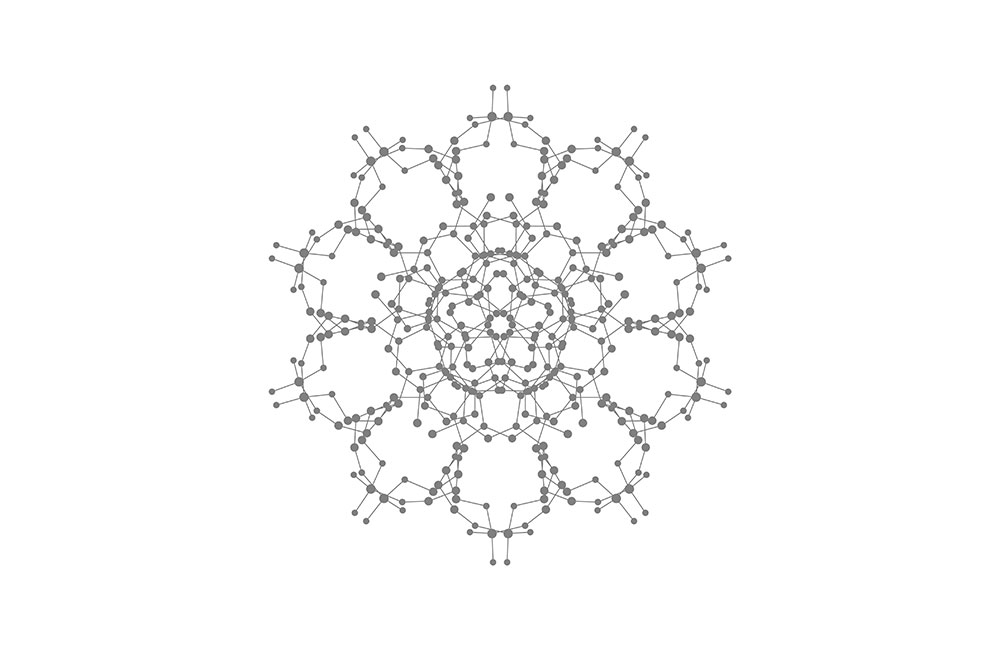
Top diagram: a DNA molecule looked through the direction of double helix.
Top: Schematic of DNA double helix.
Bottom: DNA atomic model (hydrogen atoms not shown).
The structure of DNA. In 1953, Watson and Crick published the famous double helix structure of DNA and revealed its replication mechanism. This has been deemed as one of the most important events in the history of life science, and also marked the inception of modern human genomics. In 1962, Watson, Crick and Wilkins won the Nobel Prize in physiology or medicine. In April 2003, scientists completed the human genome project by decoding 3 billion base pairs. More than 20,000 genes that encode proteins are discovered but this amount only accounts for 1.5% of total DNA. The rest genes are called “non-coding” DNA, whose function is still unknown. In September 2003, the National Human Genome Research Institute started the project of the encyclopedia of DNA elements (referred as Encode) to find out all functional components in human genome and establish their relationship to human health. The Encode project announced in 2012 that at least 80% non-coding DNA possessed one or more than one types of biological activity, and many of them related to the gene expression. The Encode project had scientists realize that the regulation of gene express was much more complicated in practice than of previous theories. The understanding to our own genetic codes is still in a rapid development. [Figure reference: Watson, J. D. and Crick F. H. Nature 171, 737 (1953); The atomic coordinates of DNA are generated by w3DNA.]
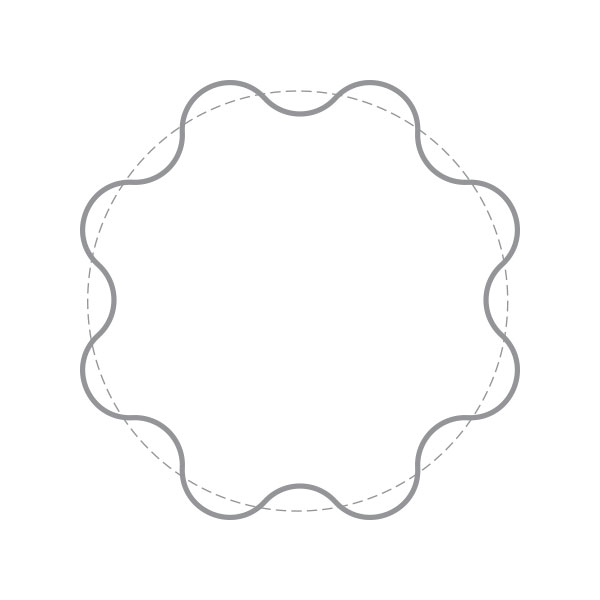
The 3D structure of myoglobin. In 1958, Kendrew et al. published the structure of myoglobin, the first complex structure of proteins by the means of X-ray diffraction analysis. Myoglobin is a protein that stores oxygen in human body and animals. Kendrew analyzed 2600 atoms in myoglobin and the structural analysis of such a complex structure was actually a great victory in science. It has taken more than 20 years for many scientists to solve this structure. At a primary stage of computer graphics, it was a great challenge to show such a structure to public. In 1961, Kendrew was invited by Scientific American to write an article regarding the three-dimensional structure of myoglobin. Geis, who had received training as architecture, spent six months on creating a spectacular illustration of the structure of myoglobin. The above image is the computer graphic created by the author to imitate Geis’s original drawing of the myoglobin structure in 1961, in order to salute to his pioneering work in the field of molecular visualization. [Figure reference: Kendrew, J. C. Sci. Am. 205, 96 (1961); Watson, H. C. Prog. Stereochem. 4, 299 (1969)]
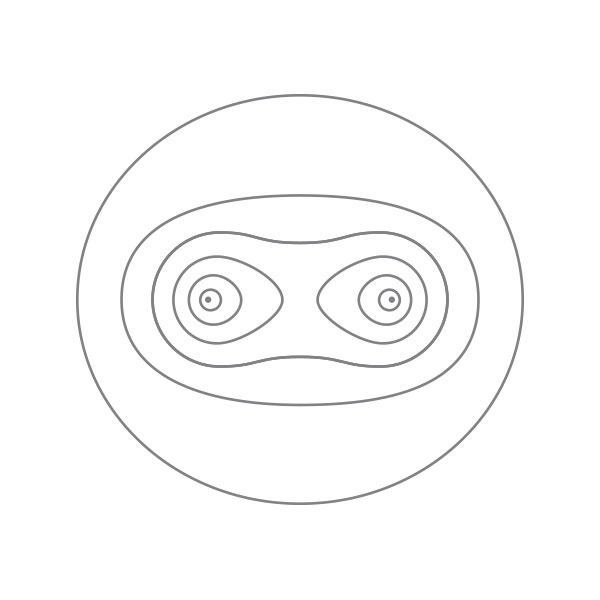
The 3D structure of hemoglobin. In 1960, Perutz published the three-dimensional structure of hemoglobin. This was the second identified complex protein structure by X-ray diffraction after Kendrew’s myoglobin. Hemoglobin contains four similar protein subunits to myoglobin. The main function of hemoglobin is to transport oxygen in blood. The critical step to solve such structure was to adopt the substitution method of heavy atom that was published by Kendrew in 1954. It took Perutz six more years to obtain the final structure of hemoglobin. After the determination of the hemoglobin structure, Perutz prioritized his research on this protein. One of his important contributions is to explain the mechanism of oxygen combination and releasing of hemoglobin at a molecular level. In 1962, Perutz shared the Nobel Prize in chemistry because of his pioneering work on the structural analysis of hemoglobin. [Figure reference: Dickerson, R. E. and Geis, I. Hemoglobin (1983)]
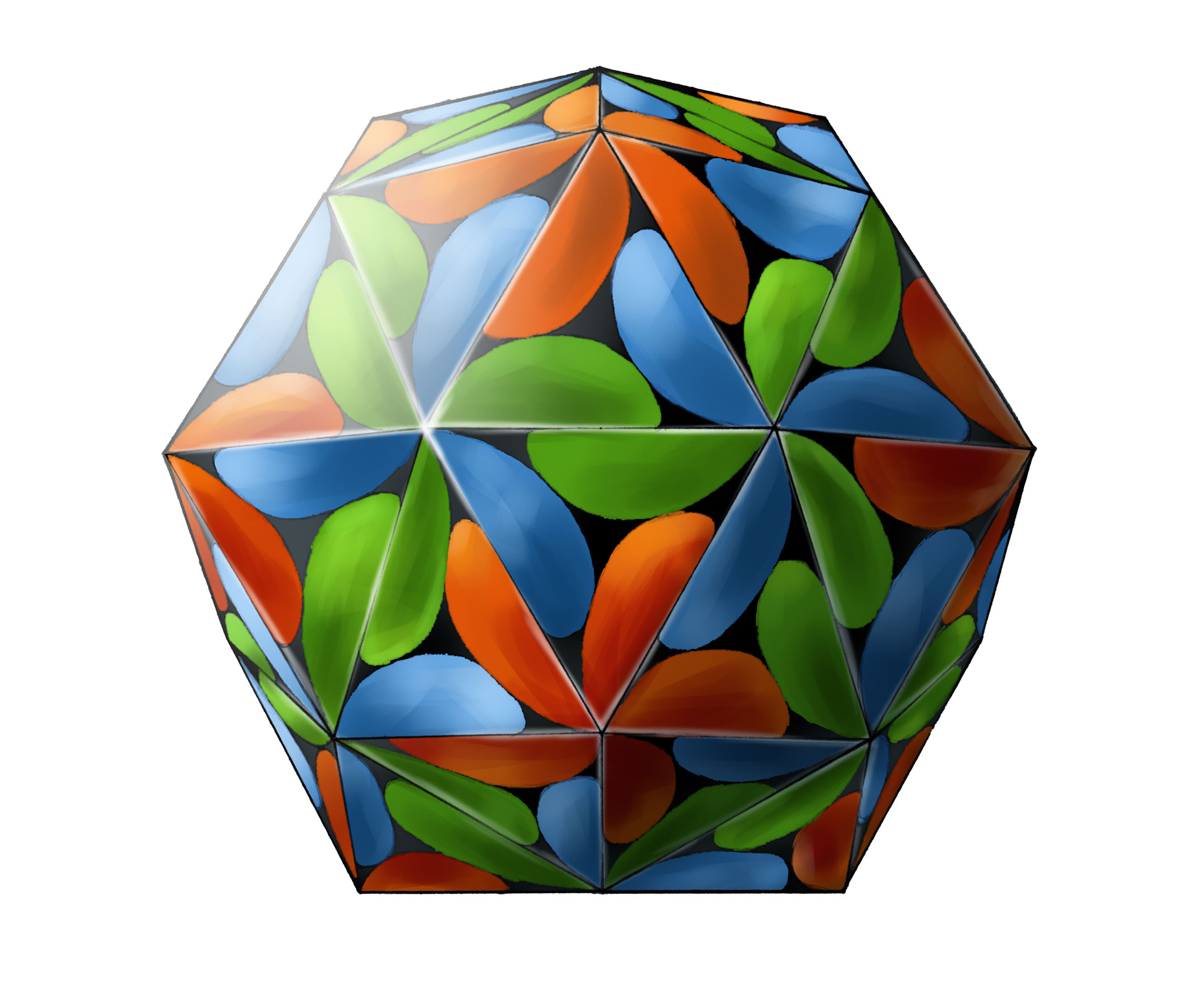
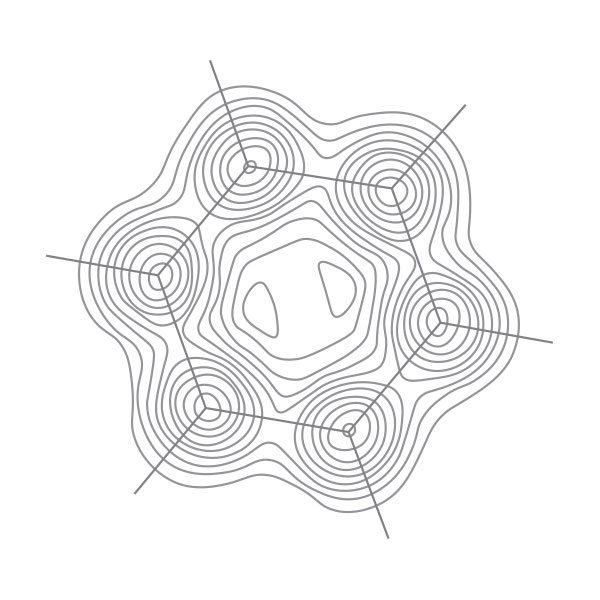
Tomato bushy stunt virus. After myoglobin and hemoglobin, more protein structures were determined. By 1976, a total of 31 protein structures were solved. The three-dimensional structure of the capsid of tomato bushy stunt virus, reported by Harrison et al. in 1978, raised the complexity of known protein structures to a new leve. The capsid of tomato bushy stunt virus contains 180 protein subunits and its spherical structure shows icosahedral symmetry. Three different types of proteins are contained in 180 subunits and represented by red, green, and blue colors, respectively. 120 protein subunits may interact with the RNA of the virus, which is enclosed in the capsid in the process of virus self-assembly. Later on, structural biologists determined more complex structures of proteins with critical physiological functions, for instance, the reaction center of photosynthesis (1984), ATP synthetase (1994), ribosome (2000), and spliceosome (2015). All these determined structures have contributed to the understanding the mysteries of life at a molecular level. The above image was adopted from Geis’s original work in 1984. The capsid structure of tomato bushy stunt virus, which was generated with atomic coordinates by using computer-graphics software can be found here. [Figure reference: HHMI Bulleting, April 2000].
Other Topics